{{
}}
Citizen science & machine learning for plant Identification: challenges from Pl@ntNet
Joseph Salmon
IMAG, Univ Montpellier, CNRS, Inria, Montpellier, France
Consortium Pl@ntnet
Pl@ntNet insights
Pl@ntNet: ML for citizen science
A citizen science platform using machine learning to help people identify plants with their mobile phones

Alexis Joly
Primary investigator, INRIA
ResearchGate

Pierre Bonnet
Primary investigator, CIRAD
ResearchGate

Hervé Goëau
Researcher, CIRAD
ResearchGate

Antoine Affouard
Backend & Staff engineer, INRIA
LinkedIn

Jean-Christophe Lombardo
IA engineer, INRIA
LinkedIn

Mathias Chouet
Backend engineer, CIRAD
GitHub

Thomas Paillot
Front engineer, INRIA
LinkedIn

Rémi Palard
Geo & Fullstack engineer, CIRAD
LinkedIn

Vanessa Hequet
Botanist, IRD
LinkedIn

Murielle Simo-Droissart
Botanist, IRD
ResearchGate

Théo Simoes
Backend engineer, INRAE
LinkedIn

Jean-Marc Sadaillan
Project manager, INRAE
LinkedIn

Christophe Botella
Researcher, INRIA
ResearchGate

Joseph Salmon
Researcher, INRIA
Website

Benjamin Bourel
Researcher, INRIA
Website

Théo Larcher
PhD candidate, INRIA
LinkedIn

Giulio Martellucci
PhD candidate, INRIA
LinkedIn

Raphaël Benerradi
PhD candidate, INRIA
LinkedIn

Ilyass Moummad
Post-doc, INRIA
Personal website | Google Scholar

Note: I am mostly innocent, I started working with the Pl@ntNet team in 2020
- Collaborative effort, involving:
- machine learners
- ecologist
- engineers
- amateurs
- Open problems:
- theoretical
- methodological
- computational (due to the size of the problems)
- interfacing many disciplines
We need you: come and help us improve it!
Contributions
- Pl@ntNet-300K (Garcin et al., 2021): Creation and release of a large-scale dataset sharing the same property (Long Tail!) as Pl@ntNet; available for the community to improve learning systems

- Prediction uncertainty quantification with long tail data (Ding et al., 2026) : providing prediction sets with statistical guarantees with Conformal Prediction

- Learning & crowd-sourced data (Lefort et al., 2024) and (Lefort et al., 2025): How to leverage multiple labels per image to improve the model? Need to assert quality: the workers, the images/labels, the model, etc.

Pl@ntNet data characteristics
_version_2.png) @ Rkitko (Wikimedia Commons)
@ Rkitko (Wikimedia Commons)
 RBG Kew
RBG KewCC-BY-SA
Intra-class variability
 Benoît Janichon
Benoît JanichonCC-BY-SA Guizotia abyssinica (L.f.) Cass.
 Patrice SIROT
Patrice SIROTCC-BY-SA Diascia rigescens E.Mey. ex Benth.
 Borquez Vicent
Borquez VicentCC-BY-SA Lapageria rosea Ruiz & Pav.
 Наталья
НатальяCC-BY-SA Casuarina cunninghamiana Miq. LC
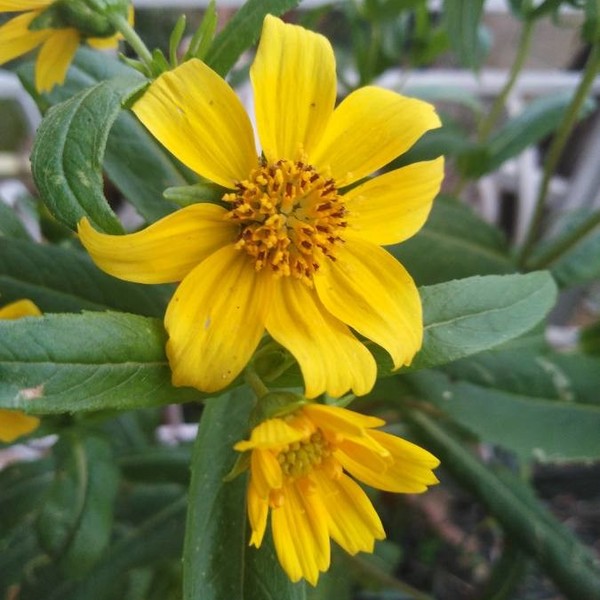 Annette Bejany
Annette BejanyCC-BY-SA Guizotia abyssinica (L.f.) Cass.
 A Lee
A LeeCC-BY-SA Diascia rigescens E.Mey. ex Benth.
 Daniel Barthelemy
Daniel BarthelemyCC-BY-SA Lapageria rosea Ruiz & Pav.
 Campos Ignacio
Campos IgnacioCC-BY-SA Casuarina cunninghamiana Miq. LC
Inter-class ambiguity
 stefano mazzotti
stefano mazzottiCC-BY-SA Cirsium rivulare (Jacq.) All.
 buqa Jarmil
buqa JarmilCC-BY-SA Chaerophyllum aromaticum L.
 Walter Reider
Walter ReiderCC-BY-SA Adenostyles leucophylla Rchb.
 furs
fursCC-BY-SA Petrosedum montanum (Songeon & E.P.Perrier) Grulich
 Rene Weck
Rene WeckCC-BY-SA Cirsium tuberosum (L.) All.
 Jcm Arthur
Jcm ArthurCC-BY-SA Chaerophyllum temulum L.
 pierre Lamy
pierre LamyCC-BY-SA Adenostyles alpina (L.) Bluff & Fingerh.
 Wolfi 41
Wolfi 41CC-BY-SA Petrosedum rupestre (L.) P.V.Heath
Sampling bias
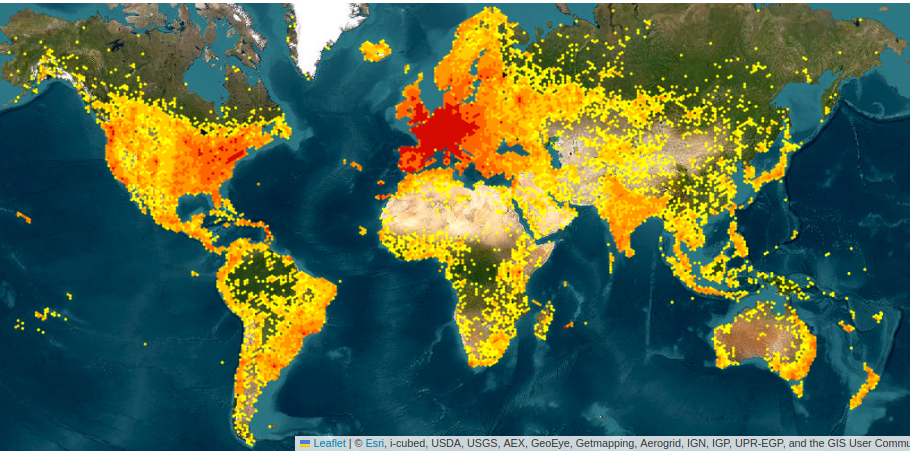
CC-BY-SA
CC-BY-SA
CC-BY-SA
CC-BY-SA
CC-BY-SA
 Dieter Wagner
Dieter WagnerCC-BY-SA
 David Eickhoff − EOL
David Eickhoff − EOLCC-BY-SA
 Patrick Cartier
Patrick CartierCC-BY-SA
 Maximilien Perrin
Maximilien PerrinCC-BY-SA
Many more biases …
- Selection bias
- Convenience sampling: easily vs. hardly accessible
- Preference for certain species: visibility / ease of identification
- Subjective bias: selection based on personal judgment, may not be random or representative
- Rare species: rare or endangered species may be under-represented
- Temporal bias / seasonal variation: seasonal changes in plant characteristics
- …
The Pl@ntNet-300K dataset
Popular datasets limitations:
- structure of labels too simplistic (CIFAR-10, CIFAR-100)
- tasks too easy to discriminate (MNIST)
- too well-balanced, with same number of images per class (Imagenet)
- contains duplicate, low-quality, or irrelevant images (Recht et al., 2019)
Release a dataset sharing similar features as the Pl@ntNet dataset to foster research in plant identification
\(\implies\) Pl@ntNet-300K (Garcin et al., 2021)
“The collective behavior induced by frictionless research exchange is the emergent superpower driving many events that are so striking today.” (Donoho, 2024)
Construction of Pl@ntNet-300K
- Earth: 400K+ species
- Pl@ntNet: 80K+ species
- Pl@ntNet-300K: 1K+ species
Note: long tail preserved by genera subsampling
Pl@ntNet300K: Long tail visualization
Characteristics:
- 306,146 color images
- Size: 32 GB
- Labels: 1 000+ species
- Required 2 000 000 volunteers
Zenodo, 1 click download
https://zenodo.org/record/5645731
Code to train models
https://github.com/plantnet/PlantNet-300KPrediction
&
uncertainty quantification

Tiffany Ding
UC Berkeley
within

Jean-Baptiste Fermanian
Inria
“Conformal Prediction for Long-Tailed Classification”
T. Ding, J.-B. Fermanian and J. Salmon
ICLR 2026
Pl@ntNet: set prediction (recommendation)
Elements to help guide the users
- provide a set of possible species/labels
- display similar images from proposed species
- give a score of confidence
For an input image \(X\), propose the most probable classes \(y\) with confidence level \(1-\alpha\) (with small \(\alpha\))
\[ \mathcal{C}_{\alpha}(X) = \big\{ y : s(X,y) \geq t_\alpha \big\} \]
Conformal prediction: sets \(t_{\alpha}\) as the \((1-\alpha)\) quantile of the scores on a calibration set
Marginal coverage targets:
\[\mathbb{P}\big[ Y \in \mathcal{C}_{\alpha}(X) \big ] \geq 1 - \alpha.\]
Class conditional coverage targets:
\[\forall y,\quad \mathbb{P}\big[ Y \in \mathcal{C}_\alpha(X) | Y=y \big ] \geq 1 - \alpha.\]
The optimal set of minimum size and marginal coverage of at least \(1-\alpha\) is: \[ \mathcal{C}_{\alpha}(x) = \left\{ y : p(y|x) \geq t_\alpha \right\} \]
The optimal set of minimum size and conditional coverage of at least \(1-\alpha\) is: \[ \mathcal{C}_{\alpha}(x) = \left\{ y : p(y|x) \geq t_\alpha^{y} \right\} \]
Marginal:
calibrate \(t_{\alpha}\) on whole calibration set \((X_i, Y_i)_{i=1}^n\)
Conditional:
calibrate \(t_{\alpha}^y\) only on \((X_i, Y_i)\) such that \(Y_i = y\)
Interactive optimal set visualization
Class conditional: useless for long-tail
Targeting Macro-Coverage
\[ \text{MacroCoverage} = \frac{1}{|\mathcal{Y}|} \sum_{y \in \mathcal{Y}} \mathbb{P}\big( y \in \mathcal{C}(X) \, | \, Y = y \big) \]
The optimal set of minimum size and Macro-Coverage of at least \(1-\alpha\) is: \[ \mathcal{C}_{\alpha}(x) = \left\{ y : \frac{p(y|x)}{p(y)} \geq t_\alpha \right\} \]
Interactive optimal set visualization (II)
Experiments on Pl@ntNet-300K
Generalization : weighted Macro-Coverage
Given user-chosen class weights \(\omega(y)\) for \(y \in \mathcal{Y}\) that sum to one, we can define the \(\omega\)-weighted macro-coverage as
\[ \begin{align} \mathrm{MacroCov}_{\omega}(\mathcal{C}) = \sum_{y \in \mathcal{Y}} \omega(y) \mathbb{P}(Y \in \mathcal{C}(X) \mid Y = y). \end{align} \]
The optimal set of minimum size and Macro-Coverage of at least \(1-\alpha\) is: \[ \begin{align} \mathcal{C}^*(x) = \left\{ y \in \mathcal{Y} : \omega(y) \dfrac{p(y|x)}{p(y)} \geq t\right\}, \end{align} \]
Experiments on Pl@ntNet-300K: endangered species
\[ \omega(y) = \begin{cases} \frac{\gamma}{W} & \text{if } y \in \mathcal{Y}_{\text{at-risk}} \quad (\text{with } W = \gamma|\mathcal{Y}_{\text{at-risk}}| + |\mathcal{Y} \setminus \mathcal{Y}_{\text{at-risk}}|)\\ \frac{1}{W} & \text{otherwise}, \end{cases} \]
Crowd-sourced data:
Votes, labels & aggregation

Tanguy Lefort
Now at Seenovate
within

Benjamin Charlier
Inrae
“Cooperative learning of Pl@ntNet’s Artificial Intelligence algorithm:
how does it work and how can we improve it?”
T. Lefort et al.
Methods in Ecology and Evolution, 2025
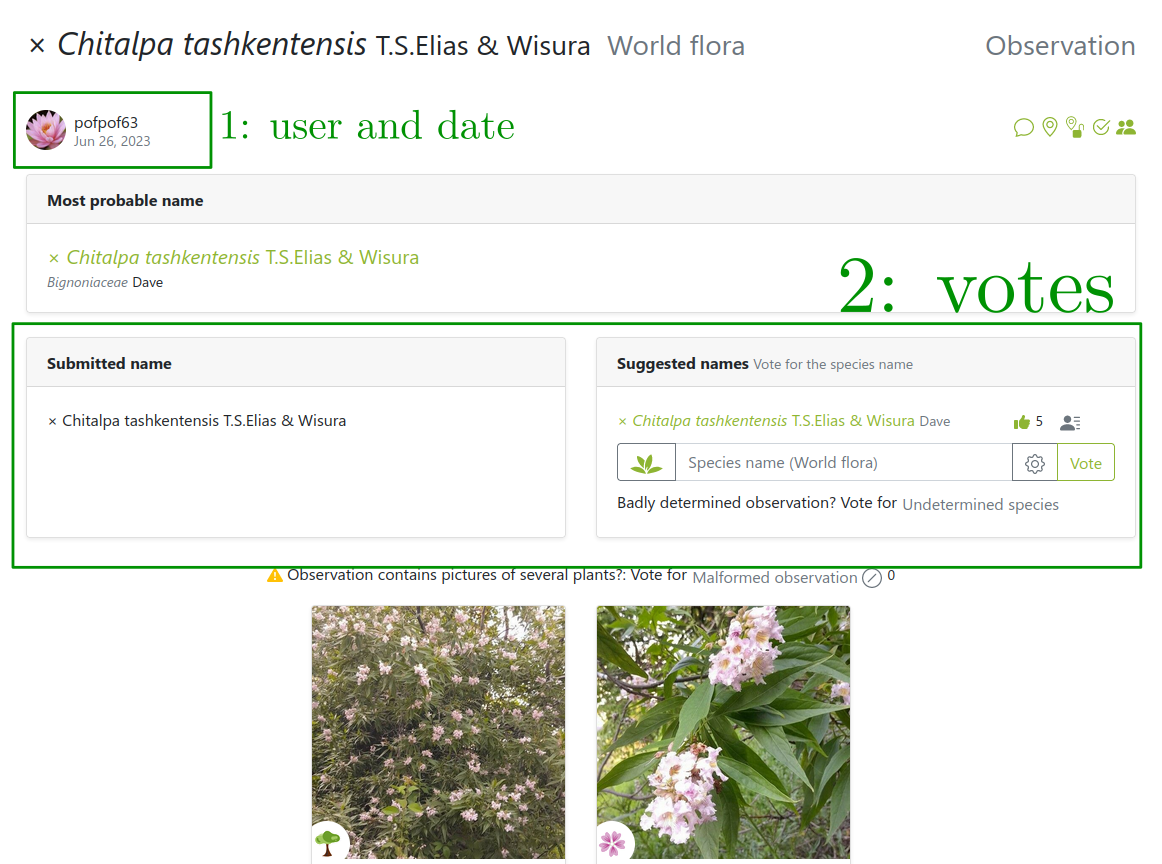
Images from users… so are the labels!
But users can be wrong or not experts
Several labels can be available per image!
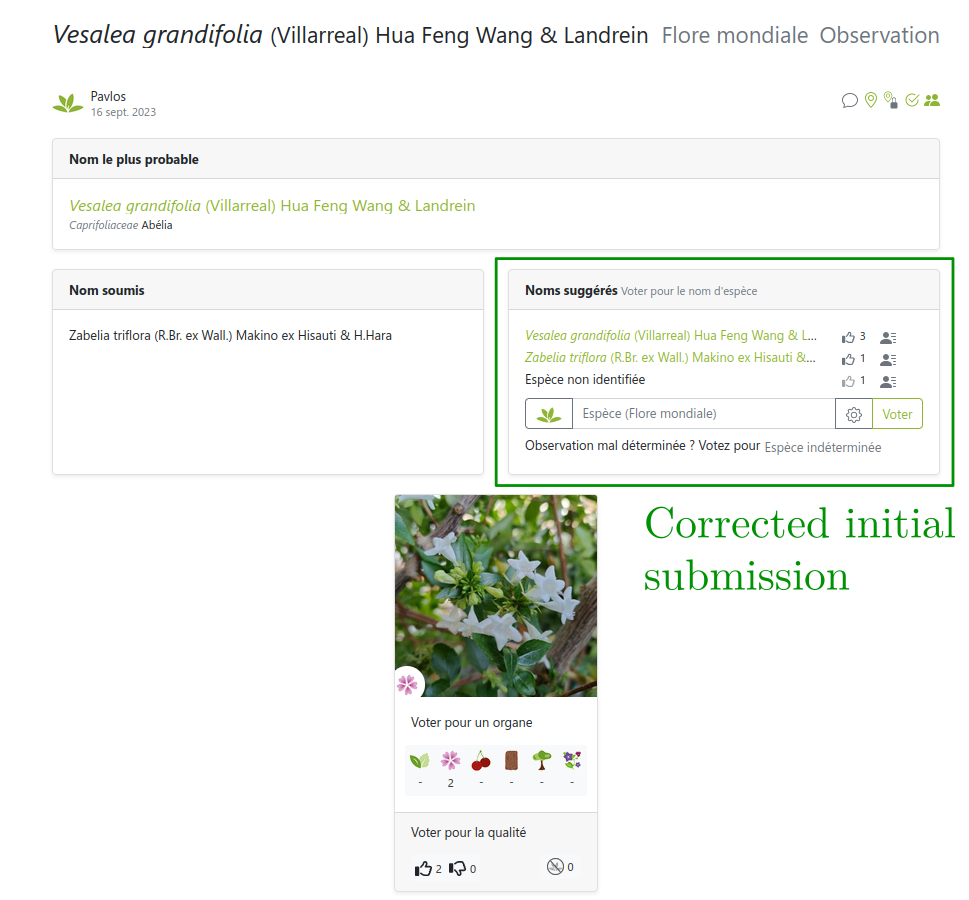
… sometimes can’t be trusted
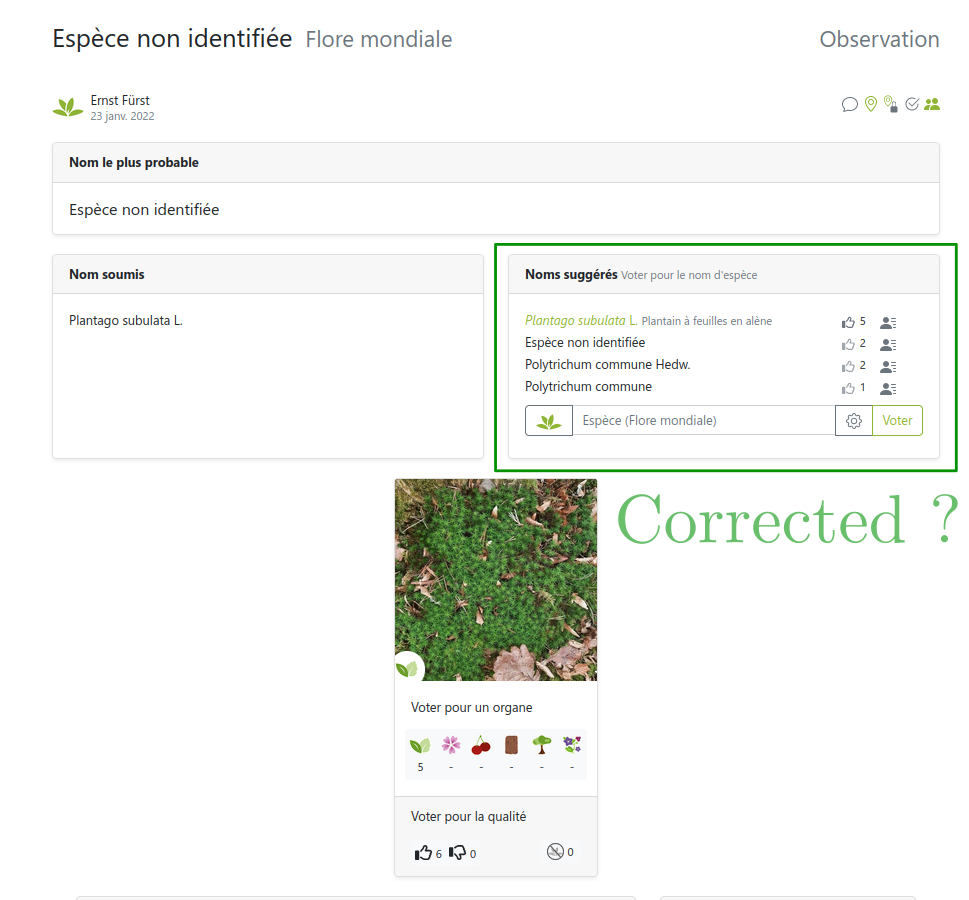
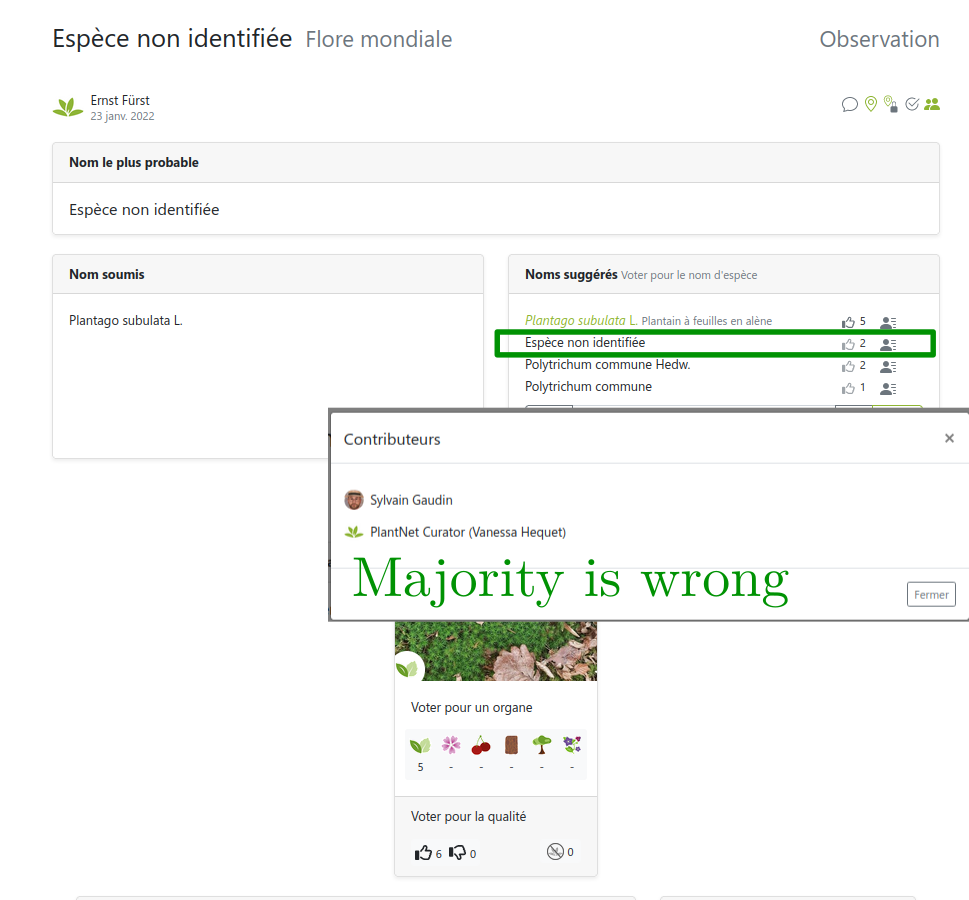
Link: https://identify.plantnet.org/weurope/observations/1012500059
Pl@ntNet label aggregation (EM algorithm)
Weighting scheme: weight user vote by its number of identified species
Take home message
- Challenges in citizen science: many and varied (need more attention)
- Crowdsourcing / Label uncertainty: helpful for data curation
- Improved data quality \(\implies\) improved learning performance
- Prediction: theory can guide the set to display
Dataset release:
- Pl@ntNet-300K: https://zenodo.org/record/5645731
- Pl@ntNet SWE flora: https://zenodo.org/records/10782465
Code release:
- Toolbox: https://peerannot.github.io/
- Some benchmarks: https://benchopt.github.io/
Future work
- Handling label hierarchy
- Human–computer interaction / performative learning / model collapse
- Improve robustness to adversarial users
- Leverage gamification for more quality labels theplantgame.com

















